As the human species evolved over the last six million years, our resident microbes did the same, adapting to vastly different conditions on our skin and in our mouths, noses, genitalia and guts.
A team of Duke University scientists has tracked how this microbial evolution unfolded, using mathematical tools originally developed for geologists.
The scientists identified microbes that diverged into new species as they colonized one area of the body after another. Their study provides a new way of looking at complicated microbial data to tease out the evolution of bacteria associated with our bodies.
The research, published in the open access journal eLife, could prompt new theories and treatments for managing these bacterial communities, collectively known as the human microbiome, to improve our personal health.
“Over the last decade, there has been significant interest in developing probiotics and transplants of beneficial bacteria to treat a wide variety of health issues,” said Lawrence A. David, Ph.D., senior author of the study and assistant professor of molecular genetics and microbiology at Duke University School of Medicine. “Our analysis gives us a window into how different bacteria adapt and evolve so that we can more effectively predict which implanted species will survive to make an impact on disease.”
Only recently have scientists begun to appreciate just how much our health depends on the trillions of bacteria that call our bodies home. We now know that these bacteria help digest the food we eat, boost our brain function, and regulate our immune systems. But figuring out how our bacteria -- which by some accounts outnumber our own cells by ten to one -- evolved to live with each other and with us has proven particularly challenging.
Scientists typically glean information on the microbiome by sampling a few million bacteria -- say, from the gut or tonsils -- and sequencing them to count which bacteria belong to each species. Then they compare those counts, generating values that tell them the relative abundance of each type of bug. But relative abundance data requires statistical methods that take into account how shifts in one species might affect another.
Justin Silverman, an MD-PhD student in the David laboratory, searched the literature for possible workarounds, and found one in an unlikely place -- the field of geology. To make sense of the relative amounts of different elements like calcium and aluminum found in rocks, geologists had devised a mathematical tool called the PhILR transform. Silverman adapted this tool to study the relative amounts of bacteria found in the microbiome.
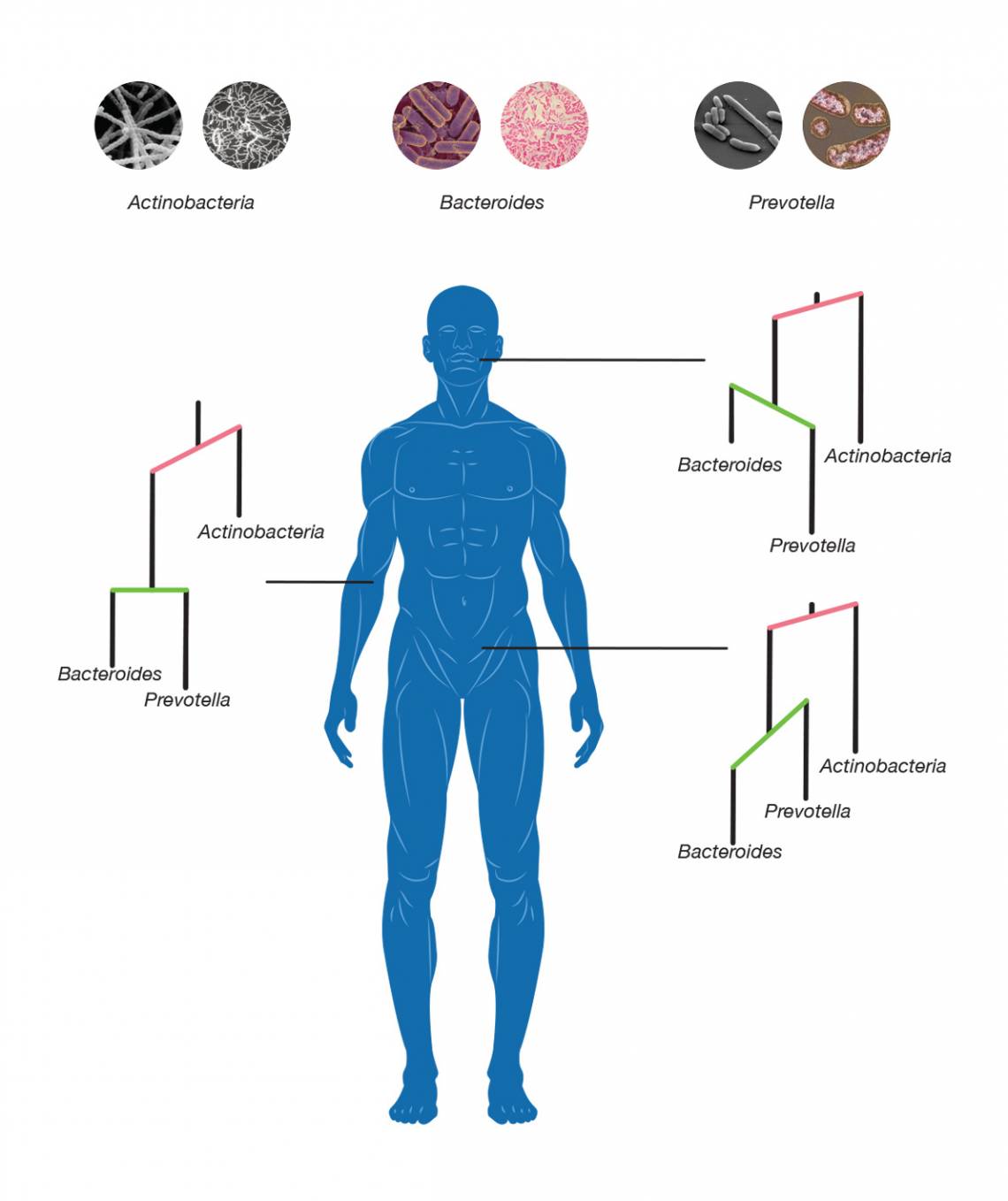
The new technique combined the sequencing counts for each species with information on their position on the bacterial family tree. The resulting statistical framework looks like a mobile you might find hanging over a baby’s crib, with a common ancestor at the top and all the subsequent generations suspended underneath, connected by a series of cross-bars. By looking at how these cross-bars tilted and swayed with the weight of the various species dangling from their tips, Silverman and his colleagues could assess how microbial communities grew and evolved in different body sites.
“This technique unlocks a tremendous toolbox of statistical methods that wouldn’t have worked before, but that can now be used to analyze microbiome data,” Silverman said.
Silverman used this framework to look at data from the Human Microbiome Project and found that different microbes have evolved to adapt to environments like our skin and mouth. For example, they discovered a group of streptococci bacteria that diverged fairly recently in different regions in the oral cavity. Our palate, tongue, throat, tonsils, gums -- even the plaque on our teeth -- each house their own species of bacteria. Such findings could help researchers determine how different genes allow microbes to adapt to one place or another, and could one day lead to new therapies that shape the microbiome.
The researchers believe their technique could be applied in practically any situation where high-throughput technologies are used to measure the composition of a sample, from the genetic makeup of a tumor to the strains of an influenza virus.
This work was supported by the Duke University Medical Scientist Training Program (GM007171), Young Investigator Grant for Probiotics Research, Searle Scholars Program and Alfred P. Sloan Foundation.
CITATION: "A phylogenetic transform enhances analysis of compositional microbiota data," Justin D. Silverman, Alex D. Washburne, Sayan Mukherjee, and Lawrence A. David. eLife, February 15, 2017. http://dx.doi.org/10.7554/eLife.21887