
Scientists can now watch how hundreds of individual cells work together to maintain and regenerate skin tissue, thanks to a genetically engineered line of technicolor zebrafish.
Every cell on the surface of the fish, from the center of the eye to the tip of each scale, is genetically programmed to glow with a slightly different hue. But these zebrafish weren’t bred to brighten up an aquarium; the colors effectively stamp each cell with a permanent barcode, letting scientists track its movements in a live animal for days or even weeks at a time.
“Before we can fully understand tissue regeneration, we need to be able to monitor what individual cells are doing,” said Kenneth D. Poss, James B. Duke Professor of Cell Biology at Duke University and lead scientist on the team that designed the color-coding system. “This is a cutting-edge way to visualize hundreds or thousands of cells at once in a regenerating tissue.”
The system, which the researchers call Skinbow, is described March 21 in the journal Developmental Cell.
Skinbow’s cells derive their technicolor hues from a technology called Brainbow, originally created to label individual neurons in the brain with distinct colors. Skin cells in the Skinbow zebrafish randomly express different combinations of red, green and blue fluorescent proteins, resulting in over 70 hues that can be reliably distinguished under a microscope. These colors are only expressed in the outermost layer of skin cells, and remain permanent throughout each cell’s lifetime.
“It is like you have given each cell an individual barcode,” said Chen-Hui Chen, a postdoctoral fellow in Poss’s lab and lead author on the study. “You can precisely see how individual cells collectively behave during regeneration.”
The team also designed software to track individual cells in a series of images collected over time, allowing them to compile full “biographies” of many different cells at once.
Using this system, the researchers followed movements and changes in shape and size of hundreds of individual skin cells over three weeks of normal skin turnover. They also monitored the behaviors of skin cells after different types of injury, from minor exfoliation to complete amputation of a fin.
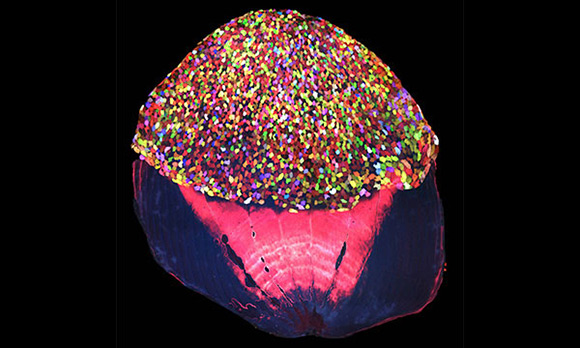
A color-coding technique called Skinbow affects the outermost layer of skin cells throughout this fish’s body. Here, the multicolor cells are evident on the exposed portion of a dissected scale. Credit: Chen-Hui Chen, Duke University
The experiments revealed a surprising diversity in cellular responses to injury. For example, following fin amputation, pre-existing cells were initially recruited from below the amputation plane to cover the wound. Then, new cells were generated at a rapid pace, and finally some cells at the tip of the regrown tissue temporarily grew in size to provide surface coverage.
“These are quite different cellular mechanisms, and one would not be able to detect the sequence or the appearance of these mechanisms without being able to track all or most of the cells on the surface of the fin,” Poss said.
In the future, the researchers hope to combine the Skinbow system with other imaging techniques to paint a more complete picture of skin tissue regeneration. It could also be applied to study how other biological stressors, such as drugs, infection or cancer, impact the cellular mechanisms underlying tissue healing.
“This is a very powerful system to see how regeneration happens,” Poss said.
Other authors include Alberto Puliafito and Luca Primo of the Candiolo Cancer Institute; and Ben D. Cox, Yi Fang and Stefano Di Talia of Duke University.
This research was supported by the National Institutes of Health (GM074057), the Finalized Research and Founding for Investments in Basic Research (RBAP11BYNP-Newton) and the National Science Foundation.
CITATION: "Multicolor Cell Barcoding for Long-Term Surveillance of Epithelial Regeneration in Zebrafish," Chen-Hui Chen, Alberto Puliafito, Ben D. Cox, Luca Primo, Yi Fang, Stefano Di Talia and Kenneth D. Poss. Developmental Cell, online March 21, 2016. DOI: 10.1016/j.devcel.2016.02.017