
Unzipping mRNA Rallies Plant Cells to Fight Infection
New tool controls protein production in living cells, with potential applications from agriculture to medicine.
Inside every cell in the body, millions of protein molecules carry out the tasks of life: They are the cellular equivalent of bricks and beams, providing structure and support. They’re also the cell’s chemical messengers, sending and receiving signals. And they’re the defenders, deployed in response to foreign invaders.
To build a protein, sections of the DNA blueprint packed inside the cell’s nucleus are transcribed into messenger molecules called mRNA, which are instructions for making proteins. These instructions are carried out to the rest of the cell, where decoding devices called ribosomes translate the mRNA’s message to assemble a chain of amino acids, the building blocks of a protein.
Normally, ribosomes scan along the mRNA molecule until they find a special three-letter sequence that says, "start here to make a protein."

But in the new study, Dong and Yezi Xiang, a Ph.D. student in Dong’s Lab, found that, when an Arabidopsis seedling detects a potential pathogen, the plant’s ribosomes bypass the usual ’start’ signal for protein synthesis and begin translating the mRNA further downstream, building a completely different chain of amino acids -- and thus a different protein -- required for fighting infection.
Dong and her team wanted to know: how do cells make the switch from one start site to another?
To better understand this rapid cellular decision-making that takes place when a plant detects an invader, the researchers turned to a technique, called SHAPE-MaP, that allows them to detect changes in mRNA folding within living cells.
Near the usual ‘green light’ that sets protein synthesis in motion, the researchers discovered short stretches of mRNA that fold back upon themselves to form double-stranded “hairpin" structures.
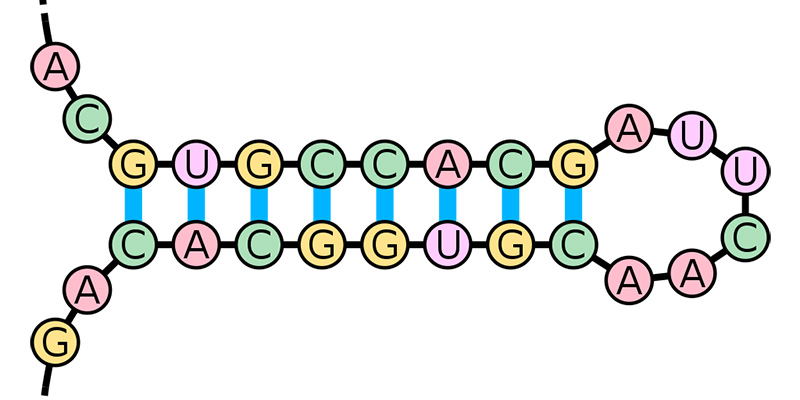
Under normal conditions, these hairpins act as brakes, preventing ribosomes from making defense proteins whose instructions lie further downstream.
But when Arabidopsis seedlings sense they’re under attack, special enzymes called RNA helicases are produced that unzip the hairpins so the ribosomes can pass through and continue scanning along the mRNA molecule.
“With these stop signs removed, the ribosomes don’t stop there, but go further down to translate defense proteins,” Dong said.
Though the team did the bulk of their experiments in Arabidopsis plants, similar RNA helicases and hairpin structures have been found in other organisms, from yeast to humans, suggesting that this mechanism for reprogramming protein synthesis may be widespread.
In follow-up experiments, the researchers used machine learning to come up with a design for a lab-made mRNA hairpin and added it to human genes. The synthetic hairpins worked to alter protein production in human cells, too.
The team has filed for a provisional patent on the discovery.
Dong says the findings could lead to new ways to engineer crops that are “not only resistant to pathogens, but also to environmental stresses like heat, cold, and drought.”
In the future, Dong said, it might also be possible to design mRNA hairpins for genome editing to help fight infections or treat diseases in people.
“The goal is to help cells produce the right amount of protein at the right time and the right place,” Dong said. “This is a step towards that goal.”
This work was supported by grants from the U.S. National Science Foundation (IOS-1645589 and IOS-2041378), the Howard Hughes Medical Institute, the State Key Research Development Program of China (2019YFA0110002), the Natural Science Foundation of China (32125007 and 91940306), and the U.S. National Institutes of Health (R35-GM122532).
Citation
"Pervasive Downstream RNA Hairpins Dynamically Dictate Start-Codon Selection," Yezi Xiang, Wenze Huang, Lianmei Tan, Tianyuan Chen, Yang He, Patrick S. Irving, Kevin M. Weeks, Qiangfeng Cliff Zhang & Xinnian Dong. Nature, Sept. 6, 2023. DOI: 10.1038/s41586-023-06500-y