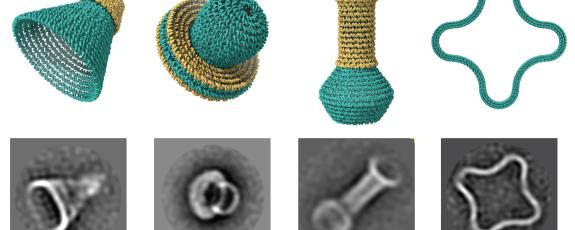
The DNA nanostructures were assembled and imaged by co-authors Raghu Pradeep Narayanan and Abhay Prasad in professor Hao Yan’s lab at Arizona State. Each tiny hollow object is no more than two millionths of an inch across. More than 50,000 of them could fit on the head of a pin.
But the researchers say these are more than mere nano-sculptures. The software could allow researchers to create tiny containers to deliver drugs, or molds for casting metal nanoparticles with specific shapes for solar cells, medical imaging and other applications.
To most people, DNA is the blueprint of life; the genetic instructions for all living things, from penguins to poplar trees. But to teams like Reif’s and Yan’s, DNA is more than a carrier of genetic information -- it’s source code and construction material.
There are four “letters,” or bases, in the genetic code of DNA, which pair up in a predictable way in our cells to form the rungs of the DNA ladder. It’s these strict base-pairing properties of DNA -- A with T, and C with G -- that the researchers have co-opted. By designing DNA strands with specific sequences, they can “program” the strands to piece themselves together into different shapes.
The method involves folding one or a few long pieces of single-stranded DNA, thousands of bases long, with help from a few hundred short DNA strands that bind to complementary sequences on the long strands and “staple” them in place.
Researchers have been experimenting with DNA as a construction material since the 1980s. The first 3D shapes were simple cubes, pyramids, soccer balls -- geometric shapes with coarse and blocky surfaces. But designing structures with curved surfaces more akin to those found in nature has been tricky. The team’s aim is to expand the range of shapes that are possible with this method.
To do that, Fu developed software called DNAxiS. The software relies on a way to build with DNA described in 2011 by Yan, who was a postdoc with Reif at Duke 20 years ago before joining the faculty at Arizona State. It works by coiling a long DNA double helix into concentric rings that stack on each other to form the contours of the object, like using coils of clay to make a pot. To make the structures stronger, the team also made it possible to reinforce them with additional layers for increased stability.
Fu shows off the variety of forms they can make: cones, gourds, clover leaf shapes. DNAxiS is the first software tool that lets users design such shapes automatically, using algorithms to determine where to place the short DNA “staples” to join the longer DNA rings together and hold the shape in place.
“If there are too few, or if they're in the wrong position, the structure won't form correctly,” Fu said. “Before our software, the curvature of the shapes made this an especially difficult problem.”
Given a model of a mushroom shape, for example, the computer spits out a list of DNA strands that would self-assemble into the right configuration. Once the strands are synthesized and mixed in a test tube, the rest takes care of itself: by heating and cooling the DNA mixture, within as little as 12 hours “it sort of magically folds up into the DNA nanostructure,” Reif said.
Practical applications of their DNA design software in the lab or clinic may still be years away, the researchers said. But “it's a big step forward in terms of automated design of novel three dimensional structures,” Reif said.
Supported By
This research was supported by the National Science Foundation (1909848, 2113941, 2004250, 1931487).
Citation
"Automated Design of 3D DNA Origami With Non-Rasterized 2D Curvature," Daniel Fu, Raghu Pradeep Narayanan, Abhay Prasad, Fei Zhang, Dewight Williams, John S. Schreck, Hao Yan, John Reif. Science Advances, Dec. 23, 2022. DOI: 10.1126/sciadv.ade4455